Harnessing HPC power with {targets}
CCT Data Science, University of Arizona
Jan 31, 2023
My Background
Ecologist turned research software engineer
Not an HPC professional or expert
Feels most comfortable never leaving the comfort of RStudio Desktop
{targets} forces you to make parallelization easy
Modular workflows & branching create independent targets
Running targets in parallel
use_targets()automatically sets things upUse
tar_make_clustermq()ortar_make_future()to run in parallelParallel processes on your computer or jobs on a computing cluster
Potentially easy entry to high performance computing
Persistent vs. Transient workers
Persistent workers with clustermq
One-time cost to set up workers
System dependency on
zeromq
Transient workers with future
Every target gets its own worker (more overhead)
No additional system dependencies
Setup clustermq on a cluster
- Take the basic HPC training at your organization
- Install
clustermqR package on the cluster - You might need to open a support ticket to get ZeroMQ (https://zeromq.org/) installed
- On the cluster, in a directory, launch R and run
targets::use_targets()
Setup clustermq on a cluster
- Edit the SLURM (or other scheduler) template that was created
#!/bin/sh
#SBATCH --job-name={{ job_name }} # job name
#SBATCH --partition=hpg-default # partition
#SBATCH --output={{ log_file | logs/workers/pipeline%j_%a.out }} # you can add .%a for array index
#SBATCH --error={{ log_file | logs/workers/pipeline%j_%a.err}} # log file
#SBATCH --mem-per-cpu={{ memory | 8GB }} # memory
#SBATCH --array=1-{{ n_jobs }} # job array
#SBATCH --cpus-per-task={{ cores | 1 }}
#SBATCH --time={{ time | 1440 }}
source /etc/profile
ulimit -v $(( 1024 * {{ memory | 8192 }} ))
module load R/4.0 #R 4.1 not working currently
module load pandoc #For rendering RMarkdown
CMQ_AUTH={{ auth }} R --no-save --no-restore -e 'clustermq:::worker("{{ master }}")'- Check that
clustermqworks - Check that
tar_make_clustermq()works
How to work comfortably
May need to run from command line without RStudio
Options for RStudio could be clunky
Data store (
_targets/) not synced with local computer
Develop local, sync, run on cluster
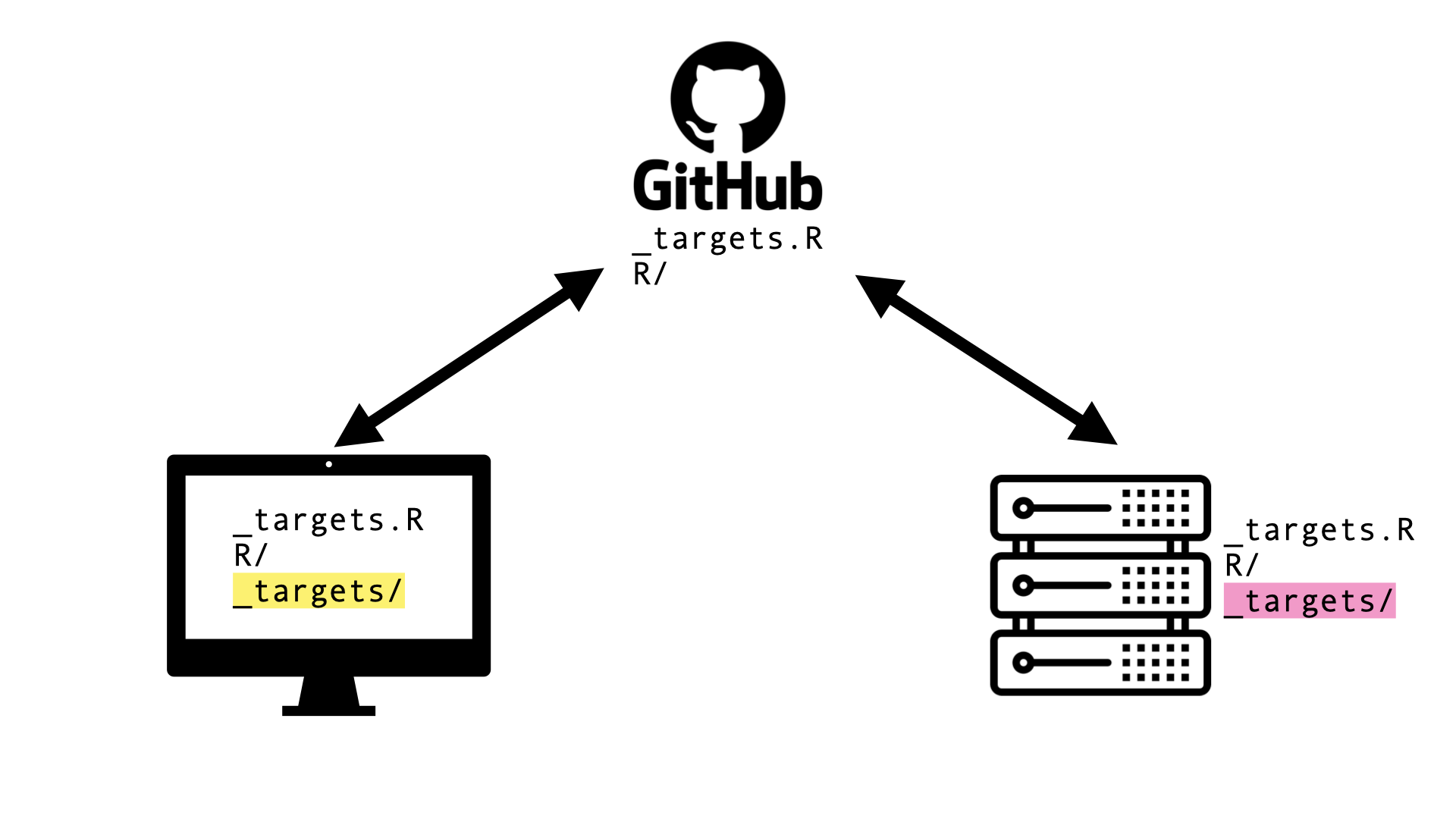
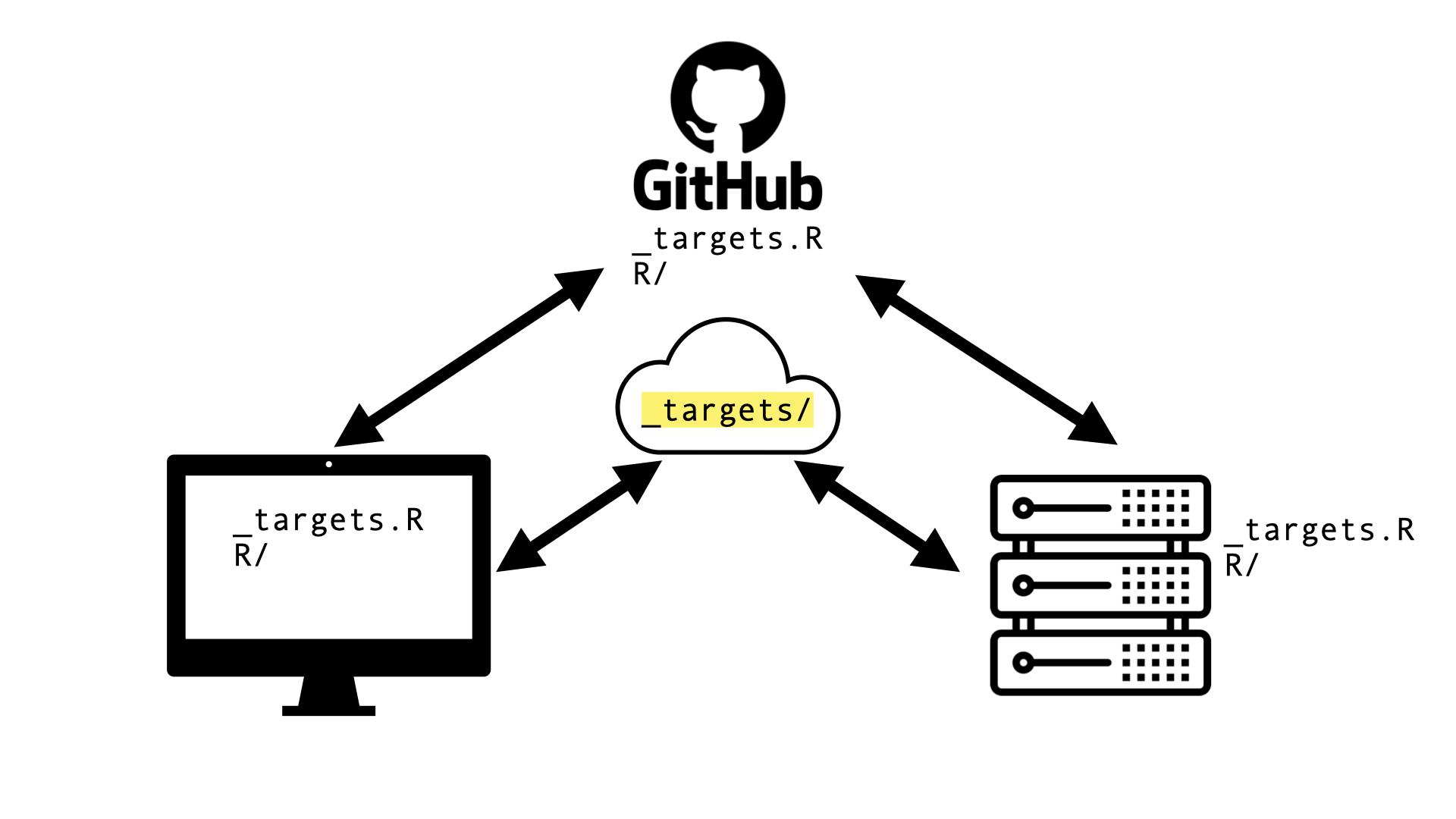
Cloud storage
SSH connection
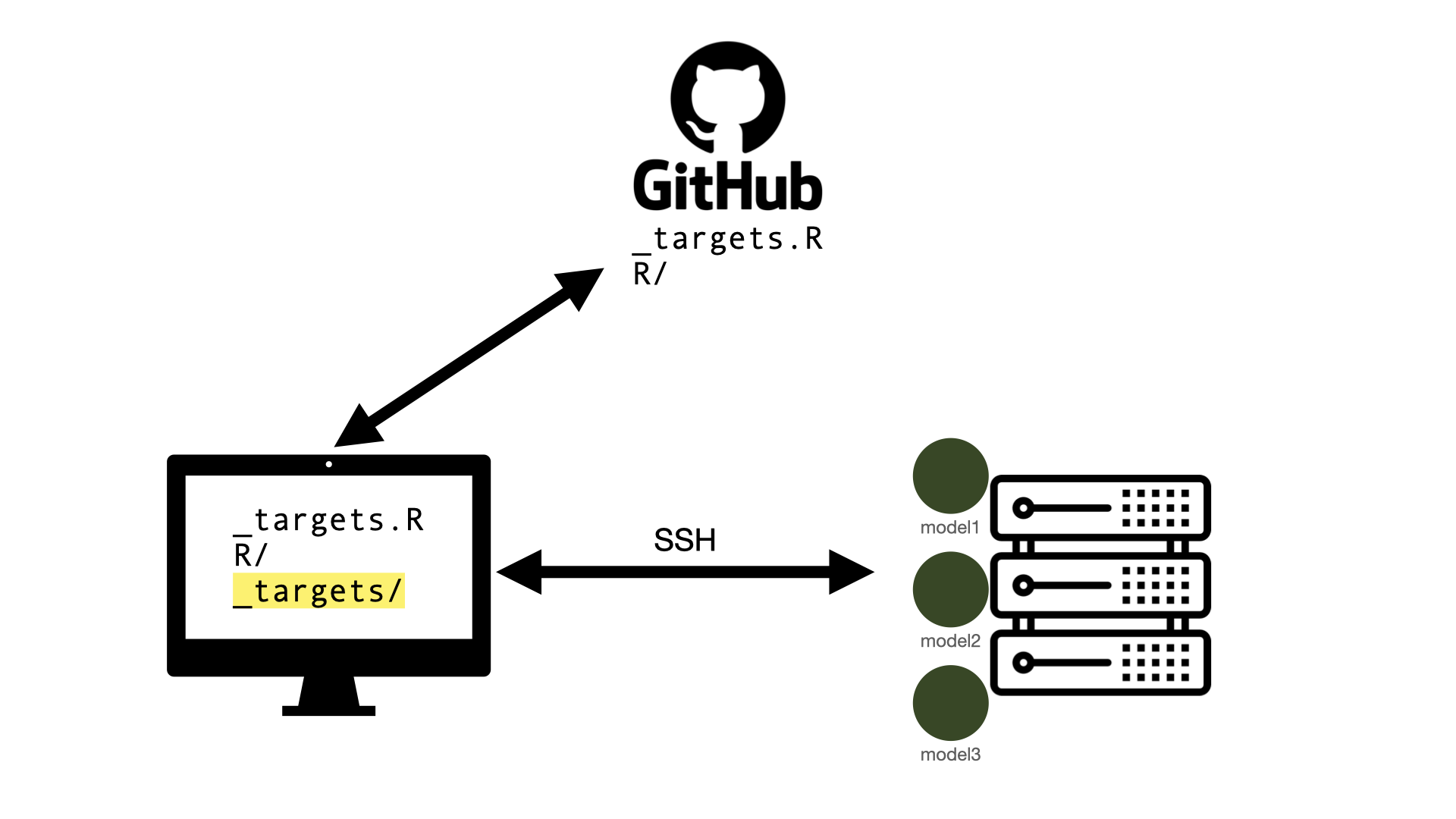
SSH connection
Develop and run workflow on your computer
Targets are sent off to the cluster to be run as SLURM jobs
Results returned and
_targets/store remains on your computerIdeal when:
Only some targets need cluster computing
Targets don’t run too long
No comfortable way to use RStudio on the cluster
SSH connection setup
- Copy SLURM template to cluster
- Edit
~/.Rprofileon the cluster:
- Set options in
_targets.Ron your computer:
Note
Packages used in the pipeline need to be installed on the cluster and local computer
Lessons Learned: UF
- Transfer of R objects back and forth is biggest bottleneck for SSH connector
- 2FA surprisingly not an issue
Lessons Learned: Tufts University
- Couldn’t get
zeromqinstalled because I couldn’t get an HPC person to email me back! futurebackend worked, but overhead was too much to be helpful
Lessons Learned: University of Arizona
- SSH connector requires an R session to run on login node—not possible at UA!
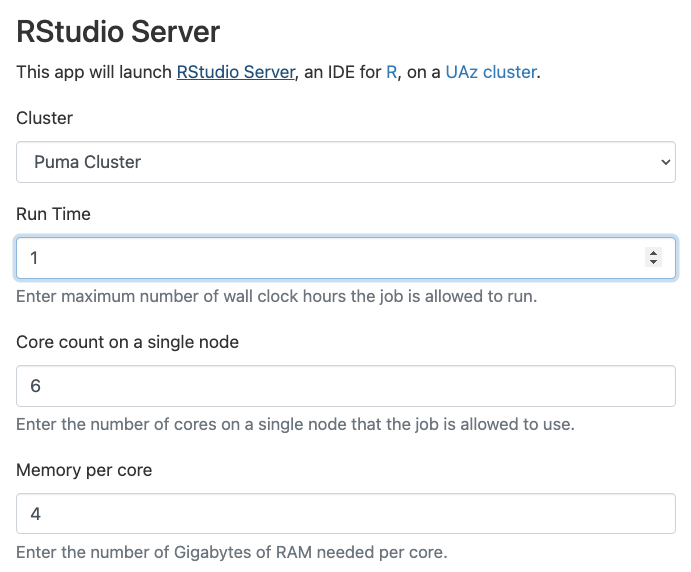
- Open On Demand RStudio Server
targetsauto-detects SLURM, but need to run as “multicore”
One last step: write it all down!
Template GitHub repo with setup instructions in README
Tell the HPC experts about it
University of Florida:
On the HPC: BrunaLab/hipergator-targets
Using SSH connector: BrunaLab/hipergator-targets-ssh
University of Arizona (WIP): cct-datascience/targets-uahpc
